Custom Query (964 matches)
Results (892 - 894 of 964)
|
Ticket
|
Owner
|
Reporter
|
Resolution
|
Summary
|
| #885 |
|
arango
|
Done
|
IMPORTANT: Update to Parallel I/O (PIO)
|
| Description |
- Several files were updated to remove compilation issues when ROMS is compiled with both PIO and ESMF/NUOPC libraries. There is a module interference with the buffer and A dummy variables in several routines inside modules. This is a weird one and a compiler bug in the ifort compiler. To avoid any issues in the future with the I/O routines, the local buffer variable is renamed to rbuffer or ibuffer for floating-point and integer representation respectively. The dummy A variable is renamed to Adat.
- Update the files in the ESM directory for WRF/WPS version 4.1 or higher. Special treatment is needed newest release of 4.3.
- Corrected a typo in set_tides.F. Many thanks to Stefan Riha for reporting this issue.
|
| #886 |
|
arango
|
Fixed
|
IMPORTANT: Corrected bug in def_state.F
|
| Description |
- The module def_state.F was missing association to def_dim.F, def_info.F, and wrt_info.F modules. We need to have:
USE def_dim_mod, ONLY : def_dim
USE def_info_mod, ONLY : def_info
...
USE wrt_info_mod, ONLY : wrt_info
The weird thing is that the linking failed when compiling with CMake and generating the shared library libroms.so. Otherwise, it was linking and running fine. It is good to know that the shared library compiling is rigorous.
- Corrected typo in get_wetdry.F. Many thanks to Ashley Brereton for bringing this issue to our attention.
|
| #887 |
|
arango
|
Done
|
Important: Enhancements and Corrections
|
| Description |
There are several corrections and enhancements in this update.
- The ecosystem header files npzd_Franks_inp.h, and npzd_Powell_inp.h, npzd_iron_inp.h where modified to read nested BioIni parameters from their respective input configuration script. Now, we have:
#ifdef ANA_BIOLOGY
CASE ('BioIni(iNO3_)')
Npts=load_r(Nval, Rval, Ngrids, BioIni(iNO3_,:))
CASE ('BioIni(iPhyt)')
Npts=load_r(Nval, Rval, Ngrids, BioIni(iPhyt,:))
CASE ('BioIni(iZoop)')
Npts=load_r(Nval, Rval, Ngrids, BioIni(iZoop,:))
CASE ('BioIni(iSDet)')
Npts=load_r(Nval, Rval, Ngrids, BioIni(iSDet,:))
#endif
The second argument to BioIni was changed from 1 to : because load_r was returning zeros for nested grid 2. Many thanks to Jose Alves for bringing this issue to our attention.
- Added missing module association in edit_multifile.F, routine edit_file_struct. Many thanks Chuning Wang for reporting this problem.
- Corrected typo in get_idata.F. We needed to have TIDE(ng)% instead of TIDE(ing)%. Many thanks to Christopher Roach for reporting this bug.
- Added def_mod_mod association to nl_roms.h driver when VERIFICATION is activated.
- The perl ROMS/Bin/dates script now accepts fractional (floating-point) values in addition to integers values in functions datenum and numdate.
- The routines ad_get_data.F, tl_get_data.F, and rp_get_data.F were updated to process surface pressure (Pair) from the forward forcing file in the 4D-Var algorithms when ATM_PRESS is activated.
- The nesting module nesting.F was updated to include U- and V-momentum averaging at the finer grid physical boundary in the fine2coarse3d routine. Similar treatment is added to the fine2coarse2d routine for Ubar and Vbar. Before, we only averaged the finer grid interior points and replaced such values in the coarser grid for two-way nesting applications. This update seems to minimize the problems reported by Guangyu Xu. His nested solution is behaving much better with the change to nesting.F:
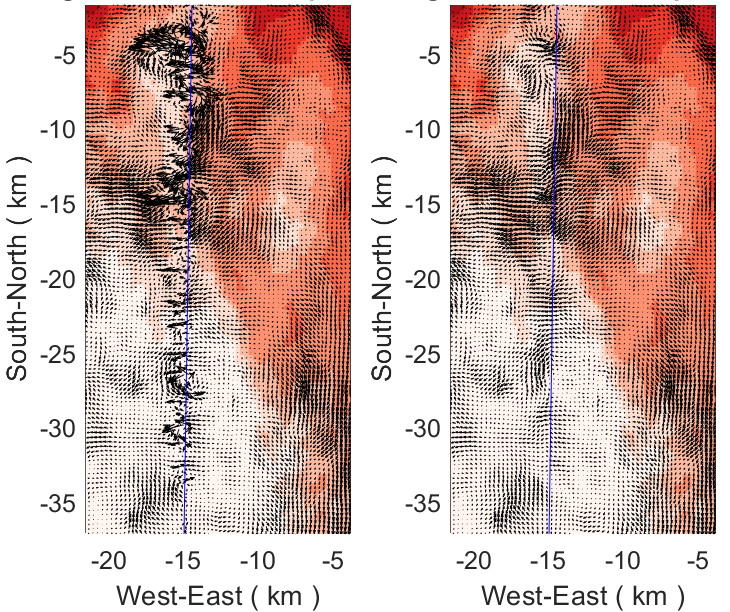
The left panel shows the velocity vector at the finer grid eastern boundary with the old version of nesting.F. The right panel shows the solution with the updated changes to nesting.F. It is quite an improvement!
Many thanks to John Warner for looking at this problem and to Guangyu Xu and Ivica Janekovic for testing this change in their applications. Ivica also reported improvements in his application.
WARNING: we will update this change to tl_nesting.F and ad_nesting.F in the near future since it will require careful testing.
|
Note:
See
TracQuery
for help on using queries.

